加州大学Vijay Ramani团队近日取得一项新成果。经过不懈努力,他们的研究开发出了单染色质纤维上普遍的和程序化的核小体畸变。该研究于2026年4月29日发表于国际一流学术期刊《自然》杂志上。
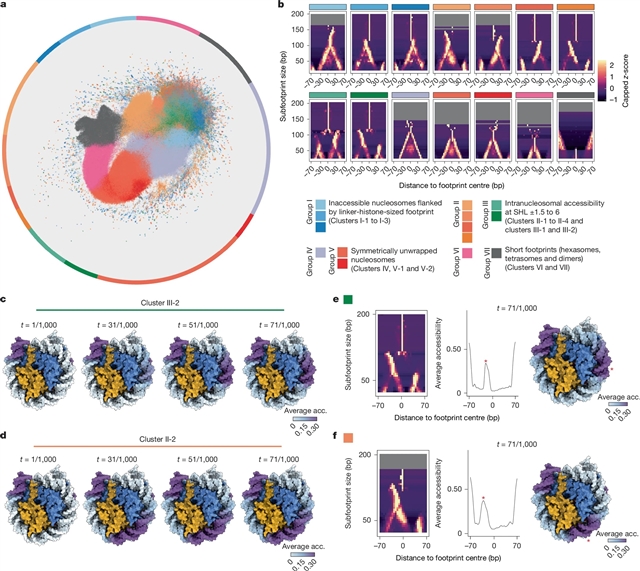
为了解决这个问题,该课题组研究人员提出了迭代定义不可接近长度(IDLI),这是一种计算方法,可以通过长读单分子足迹绘制结构不同的核小体、亚核小体和其他蛋白质-DNA相互作用的单分子共占用图。IDLI将甲基化酶在单个染色质纤维上不可接近的足迹分为(i)连接体-组蛋白相关核小体;(ii)核小体沿核小体包膜具有局灶性DNA可及性;(iii)未包裹的核小体;亚核体物种,如六体体、四体体和其他短DNA保护体。
将IDLI应用于单主题胚胎干细胞的染色质,研究组发现超过85%的核小体表现出核小体内可接近的DNA(核小体“扭曲”)。课题组观察到表观基因组结构域和表达水平特异性畸变模式,包括启动子和无主旋律卫星重复序列。转录因子(TF)基序的发生与不同类型的畸变显著相关,退化实验提供了TF直接调控的证据。该课题组研究人员将IDLI应用于人诱导多能干细胞和原代母母肝细胞的体外内胚层分化。在这两种情况下,课题组研究人员观察到先锋TF FOXA2结合位点的畸变,表明畸变是发育编码的,存在于体内。最后,小鼠遗传实验表明FOXA2的核小体结合域直接影响体内核小体结构,暗示这些蛋白-核小体相互作用是扭曲的直接介质。他们的研究表明,核小体在单分子水平上存在极端但受调节的结构变异性。
此外,他们的方法提供了在生物学背景下模拟TF结合、核小体重塑和细胞类型特异性染色质调节的机会。
研究人员表示,尽管对核小体进行了数十年的生化和结构研究,但研究人员缺乏基因组尺度的方法来确定沿单个染色质纤维的核小体结构的变异性。
附:英文原文
Title: Pervasive and programmed nucleosome distortion on single chromatin fibres
Author: Yang, Marty G., Richter, Hannah J., Wang, Simai, McNally, Colin P., Moore, Camille M., Emadi, Ali, Harris, Nicole E., Dhillon, Simaron, Maresca, Michela, Pan, Huimin, Saunders, Hayden, Yang, Ruiqiao, Ostrowski, Megan S., Anderson, Erika C., de Wit, Elzo, Maher, Jacquelyn J., Fan, Yuhong, Narlikar, Geeta J., Nora, Elphge P., Willenbring, Holger, Goodarzi, Hani, Ramani, Vijay
Issue&Volume: 2026-04-29
Abstract: Despite decades of biochemical and structural studies of the nucleosome1, researchers lack genome-scale methods to determine variability in nucleosome structure along individual chromatin fibres. To address this, here we present Iteratively Defined Lengths of Inaccessibility (IDLI), a computational method that maps the single-molecule co-occupancy of structurally distinct nucleosomes, subnucleosomes and other protein–DNA interactions through long-read single-molecule footprinting2,3. IDLI classifies methylase-inaccessible footprints on individual chromatin fibres into (i) linker-histone-associated nucleosomes; (ii) nucleosomes with focal DNA accessibility along the nucleosome wrap; (iii) unwrapped nucleosomes; and (iv) subnucleosomal species such as hexasomes, tetrasomes and other short DNA protections. Applying IDLI to chromatin from mouse embryonic stem cells, we discover that more than 85% of nucleosomes exhibit intranucleosomally accessible DNA (nucleosome ‘distortion’). We observe epigenomic-domain- and expression-level-specific patterns of distortion, including at promoters and mouse satellite repeat sequences. Transcription factor (TF) motif occurrence correlates significantly with distinct types of distortion, and degron experiments provide evidence of direct regulation by TFs. We apply IDLI to in vitro endoderm differentiation in human induced pluripotent stem cells and primary mouse hepatocytes. In both cases, we observe distortion at pioneer TF FOXA2 binding sites, demonstrating that distortion is developmentally encoded and present in vivo. Finally, genetic experiments in mice show that a nucleosome-binding domain of FOXA2 directly affects nucleosome structure in vivo, implicating these protein–nucleosome interactions as direct mediators of distortion. Our work suggests extreme but regulated nucleosome structural variability at the single-molecule level. Furthermore, our approach offers opportunities to model TF binding, nucleosome remodelling and cell-type-specific chromatin regulation across biological contexts.
DOI: 10.1038/s41586-026-10418-6
Source: https://www.nature.com/articles/s41586-026-10418-6
Nature:《自然》,创刊于1869年。隶属于施普林格·自然出版集团,最新IF:69.504
官方网址:http://www.nature.com/
投稿链接:http://www.nature.com/authors/submit_manuscript.html
