美国范·安德尔学院Hong Li团队开发出Cas9甲基化敏感编辑的分子基础。相关论文于2026年4月15日发表在《自然》杂志上。
据悉,细菌CRISPR-Cas9(Cas9)核酸酶已成为广泛生物体的强大基因组操作工具。然而,它还没有充分利用基因组中普遍存在的DNA甲基化。
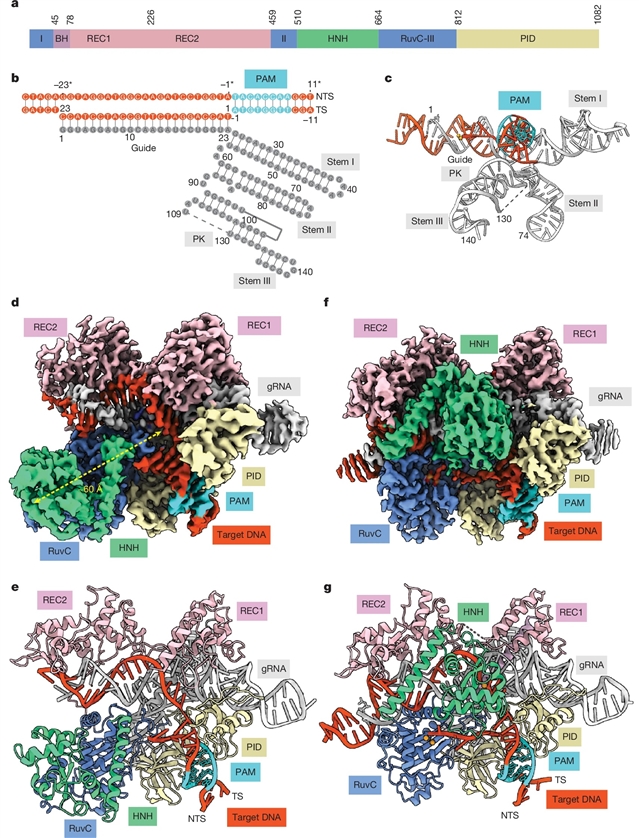
为了填补这一空白,该研究团队报告了甲基化敏感Cas9 (ThermoCas9)的生化、结构和人类基因组编辑特性。在体外,ThermoCas9能有效地结合并切割其原间隔器相邻基序(PAM) 5'-NNNNCGA-3'或5'-NNNNCCA-3'上游的DNA。然而,PAM序列(5mCpG或5mCpC)中第5个胞嘧啶的甲基化显著抑制ThermoCas9活性。研究分别以2.8 Å和2.2 Å的分辨率解析了ThermoCas9在切割前和切割后状态的冷冻电镜结构,揭示了其严格依赖未甲基化胞嘧啶进行PAM结合的分子基础,并为后续的酶工程改造提供了指导。课题组在具有不同DNA甲基化景观的人类细胞系中展示了ThermoCas9的甲基化敏感编辑。
此外,该研究团队证明了催化增强的ThermoCas9有效靶向在乳腺癌患者中持续低甲基化的腔内表达特征基因。由于其对DNA甲基化的敏感性,ThermoCas9可以特异性靶向与疾病相关的低甲基化细胞,这为基因组编辑技术增加了另一层精确度。
附:英文原文
Title: Molecular basis for methylation-sensitive editing by Cas9
Author: Roth, Mitchell O., Shu, Yuerong, Zhao, Yu, Trasanidou, Despoina, Hoffman, Renee D., Sdfeld, Christian, Bouzetos, Eugenios, Trasanidis, Nikolaos, Zawrotny, Michael, Gelasco, Mary K., Medina, Megan L., Das, Anuska, Rai, Jay, Goswami, Hemant N., Wang, Bing, van der Oost, John, Li, Hong
Issue&Volume: 2026-04-15
Abstract: The bacterial CRISPR–Cas9 (Cas9) nuclease has become a powerful genome manipulation tool for a wide range of organisms1,2,3. However, it has yet to fully leverage the pervasive presence of DNA methylation in genomes4,5,6,7,8,9,10. Here, to fill this gap, we report biochemical, structural and human genome-editing characterizations of a methylation-sensitive Cas9 (ThermoCas9). ThermoCas9 efficiently binds to and cleaves DNA upstream of its protospacer adjacent motif (PAM) 5′-NNNNCGA-3′ or 5′-NNNNCCA-3′ in vitro. Methylation of the fifth cytosine in either PAM sequence (5mCpG or 5mCpC), however, significantly inhibits ThermoCas9 activity. Cryo-electron microscopy structures of ThermoCas9 in pre-cleavage and post-cleavage states at 2.8 and 2.2 resolution, respectively, reveal the molecular basis for the stringent requirement of the unmethylated cytosine in PAM binding and provide guidance for further enzyme engineering. We demonstrate methylation-sensitive editing by ThermoCas9 in human cell lines with distinct DNA methylation landscapes. Moreover, we demonstrate that a catalytically enhanced ThermoCas9 efficiently targets luminal expression signature genes that are consistently hypomethylated in patients with breast cancer. Owing to its sensitivity to DNA methylation, ThermoCas9 can specifically target cells with disease-related hypomethylation, which adds another layer of precision to genome-editing technologies.
DOI: 10.1038/s41586-026-10384-z
Source: https://www.nature.com/articles/s41586-026-10384-z
Nature:《自然》,创刊于1869年。隶属于施普林格·自然出版集团,最新IF:69.504
官方网址:http://www.nature.com/
投稿链接:http://www.nature.com/authors/submit_manuscript.html
