Demonstration of End-to-End Automation of DNA Data Storage
期刊:Scientific Reports
作者:Christopher N. Takahashi, Bichlien H. Nguyen, Karin Strauss, Luis Ceze
发表时间:2019/03/21
论文标签:Computer science, Synthetic biology
数字识别码: 10.1038/s41598-019-41228-8
原文链接:http://t.cn/E6sz5kk
Storing information in DNA is an emerging technology with considerable potential to be the next generation storage medium of choice. Recent advances have shown storage capacity grow from hundreds of kilobytes to megabytes to hundreds of megabytes1,2,3. Although contemporary approaches are book-ended with mostly automated synthesis4 and sequencing technologies (e.g., column synthesis, array synthesis, Illumina, nanopore, etc.), significant intermediate steps remain largely manual1,2,3,5. Without complete automation in the write to store to read cycle of data storage in DNA, it is unlikely to become a viable option for applications other than extremely seldom read archival.
To demonstrate the practicality of integrating fluidics, electronics and infrastructure, and explore the challenges of full DNA storage automation, we developed the first full end-to-end automated DNA storage device. Our device is intended to act as a proof-of-concept that provides a foundation for continuous improvements, and as a first application of modules that can be used in future molecular computing research. As such, we adhered to specific design principles for the implementation: (1) maximize modularity for the sake of replication and reuse, and (2) reduce system complexity to balance cost and labor input required to setup and run the device modules.
Our resulting system has three core components that accomplish the write and read operations (Fig. 1a): an encode/decode software module, a DNA synthesis module, and a DNA preparation and sequencing module (Fig. 1b,c). It has a bench-top footprint and costs approximately $10 k USD, though careful calibration and elimination of costly sensors and actuators could reduce its cost to approximately $3 k–4 k USD at low volumes.

图1:An overview of the write-store-read process. Data is encoded, with error correction, into DNA bases, which are synthesized into physical DNA molecules and stored. When a user wishes to read the data, the stored DNA is read by a DNA sequencer into bases and the decoding software corrects any errors retrieving the original data. (a) The logical flow from bits to bases to DNA and back. (b) A block diagram representation of the system hardware’s three modules: synthesis, storage, and sequencing. (c) A photograph showing the completed system. Highlighted are the storage vessel and the nanopore loading fixture. The majority of the remaining hardware is responsible for synthesis. (d) Overview of enzymatic preparation for DNA sequencing. An arbitrary 1 kilobase “extension segment” of DNA is PCR-amplified with TAQ polymerase, and a Bsa-I restriction site is added by the primer, leaving an A-tail and a TCGC sticky end after digestion. The extension segment is then T/A ligated to the standard Oxford Nanopore Technology (ONT) LSK-108 kit sequencing adapter, creating the “extended adapter,” which ensures that sufficient bases are read for successful base calling. For sequencing, the payload hairpin and extended adapter are ligated, forming a sequence-ready construct that does not require purification.
Before a file can be written to DNA, its data must first be translated from 1’s and 0’s to A’s, C’s, T’s, and G’s. The encode software module is responsible for this translation and the addition of error correction into the payload sequence (see the Methods section and work by Richard Hamming6). Once the payload sequence is generated, additional bases are added to ensure its primary and secondary structure is compatible with the read process and the DNA sequence is sent to the synthesis module for instantiation into physical DNA molecules.
The DNA synthesis module is built around two valved manifolds that separately deliver hydrous and anhydrous reagents to the synthesis column. Our initial designs used standard valves, but the dead volume at junction points caused unacceptable contamination between cycles. Therefore, we switched to zero dead volume valves7. The combined flow path is then monitored by a flow sensor, whose output is coupled to a standard fitting; the fitting can be coupled to arbitrary devices, such as a flow cell for array synthesis8 or, in this case, adapted to fit a standard synthesis column. Once synthesis is complete, the synthesized DNA is eluted into a storage vessel, where it is stored until retrieval.
When a read operation is requested, the stored DNA pool’s volume is reduced to about 2 μL to 4 μL by discarding excess DNA through the waste port. A syringe pump in the DNA preparation and sequencing module then dispenses our single-step preparation/sequencing mix (Fig. 1d) into the storage vessel; positive pressure pushes the mixture into the ONT MinION’s priming port (Figs 1b,c). We chose the MinION as our sequencing device due to its low cost, ease of automation, and high throughput. However, it is neither capable of reading unmodified DNA, nor is it optimized for reading short DNA oligonucleotides9. In particular, we have observed that reads shorter than 750–1000 bases tend to get missed or discarded by the MinION’s software. To mitigate these limitations, we developed a single-step MinION preparation protocol that requires only payload DNA and a master mix containing a customized adapter (Fig. 1d) with a 1 kbase extension region, T4 ligase, ATP, and a buffer. Each payload sequence is constructed to form a hairpin structure with a specific 5′ 4-base overhang. The customized adapter has a complementary overhang, which aids T4-mediated, sticky-ended ligation. To sequence, the payload and master mix are combined and incubated at room temperature for 30 minutes. Thereafter, the mixture is directly loaded into the MinION through the priming port. Since the introduction of air bubbles causes sequencing failure, we built a 3D printed bubble detector that valves off the loading port immediately after detecting the gas that is aspirated following the sample. This allows the system to load nearly the full sample without damaging the flow cell. Additionally, while not demonstrated here, other research suggests that random access via selective ligation over a small set of sequence identifiers (≈20) can be achieved using orthogonal sticky ends during preparation10.
Once sequencing begins, the decode software module aligns each read to the 1 k base extension region and the poly-T hairpin. If the intervening region of DNA is the correct length, the decoder attempts to error check/correct the payload using a Hamming code with an additional parity bit; the code corrects all single-base errors and detects all double-base errors. Once the payload is successfully decoded, it is considered correct if it matches a 6-base hash stored with the data. At this point, sequencing terminates, and the MinION flow cell may be washed and stored for later reuse.
Our system’s write-to-read latency is approximately 21 h. The majority of this time is taken by synthesis, viz., approximately 305 s per base, or 8.4 h to synthesize a 99-mer payload and 12 h to cleave and deprotect the oligonucleotides at room temperature. After synthesis, preparation takes an additional 30 min, and nanopore reading and online decoding take 6 min.
Using this prototype system, we stored and subsequently retrieved the 5-byte message “HELLO” (01001000 01000101 01001100 01001100 01001111 in bits). Synthesis yielded approximately 1 mg of DNA, with approximately 4 μg ≈ 100 pmol retained for sequencing. Nanopore sequencing yielded 3469 reads, 1973 of which aligned to our adapter sequence. Of the aligned sequences, 30 had extractable payload regions. Of those, 1 was successfully decoded with a perfect payload. The remaining 29 payloads were rejected by the decoder for being irrecoverably corrupt.
Inspecting the sequencing data indicates that the low payload yield and decode rate was largely due to two factors. The first and primary factor is low ligation efficiency. Although chemical conditions should be optimal for T4 ligase, incomplete strands from the unpurified synthesis product likely out-competed full-length strands, leading to a poor apparent ligation rate of less than 10% (Fig. 2c). The second factor is read and write fidelity. To interrogate the write error rate, we synthesized a randomly generated 100-base oligonucleotide with distinct 5′ and 3′ primer sequences. The oligonucleotide was then PCR-amplified and sequenced with an Illumina NextSeq instrument to reveal: an error rate of almost zero insertions; <1% substitutions; and 1–2% deletions (Fig. 2a) for most positions, with increased deletions toward the 5′ end due to increased steric hindrance as strand length increases11. Literature suggests a nanopore error rate near 10%9,12, so we also performed a synthesis-to-sequencing error rate analysis on an 89-mer hairpin sequence, encoding “HELLO” in its first 32 payload bases. Figure 2b shows the read error when aligned to the extended adapter and payload sequence. Bases −60 to −1 were directly PCR-amplified from the lambda genome and given a good baseline for nanopore sequencing fidelity under our conditions; bases 0 through +40 come from the payload region and characterize the total write-to-read error rate. The complex combination of these errors — especially deletions and read truncations — causes many strands to be discarded before a decoding attempt is made. Indeed, of 25,592 reads in this new dataset, 286 aligned well in the −100 to −1 region (score > 400) and contained enough bases to attempt decoding. Of those 251 had uncorrectable corruption, 11 had invalid checksum bases after correction, 8 were corrupted but correctable and of those 3 had hashes in agreement, 16 were perfect reads, and 0 were decoded but contained the wrong message.
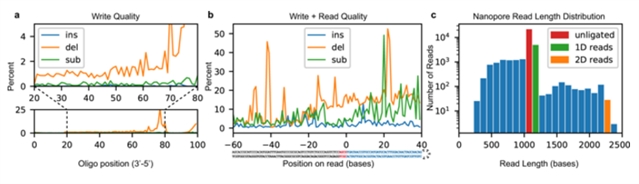
图2:Synthesis and sequencing process quality. (a) Insertion, deletion, and substitution frequency by locus for a synthesized and PCR-amplified 100-mer. Below: An overview of errors. Above: An expanded view of the central 60 bases. The terminal 20 bases come from primers used in amplification and therefore are not representative of synthesis quality. (b) Combined write-to-read quality of synthesis, ligation, and sequencing. Bases −60 to −4 (below, grey) are adapter bases. Bases −3 to 0 (below, red) are the ligation scar. Bases 0 to 39 (below, blue) are the synthesized payload region with 8 bases of padding on the 3′ end. (c) Distribution of nanopore read lengths with unligated, 1D and 2D read lengths identified.
We demonstrated the first fully automated end-to-end DNA data storage device. This device establishes a baseline from which new improvements may be made toward a device that eventually operates at a commercially viable scale and throughput. While 5 bytes in 21 hours is not yet commercially viable, there is precedent for many orders of magnitude improvement in data storage13. Infact, recent storage advances by Erlich et al.2 of 2 Mbytes and Organick et al. of 200 Mbytes3demonstrate orders of magnitude improvements in the past two years and the underlying physics and chemistry show impressive upper bounds for density3.
Furthermore, the modules and methods developed here are now being applied to other molecular computing projects internally. For example, by using a non-cleavable linker in the synthesis column and adding a reagent port for chip-synthesized DNA, we can use the same platform to perform a database query in DNA14. Additionally, our sequencing preparation protocol and loading hardware can be adapted for use with our digital microfluidics platform15 and used as a readout for DNA strand displacement reactions.
Near-term improvements will focus primarily on system optimizations in synthesis, cycle count, and cost. Synthesis time can be reduced by 10–12 hours with the addition of heat in the cleave step16. Multiple writes (with or without reads) can be achieved by the addition of additional synthesis columns and a fluid multiplexer. Multiple reads can also be achieved with minor modifications (Supplemental Section 1) and exploiting the MinION flow cell’s reusability. Additionally, a cost-optimized version could be designed by eliminating the syringe pump and flow sensor, both unnecessary if flow rates are well measured and calibrated. This could save approximately 60% of our current device’s cost at the expense of more laborious operation. Future improvements will focus on bringing storage density, coding, and sequencing yield up to parity with modern manual and semi-automated methods.
摘要:Synthetic DNA has emerged as a novel substrate to encode computer data with the potential to be orders of magnitude denser than contemporary cutting edge techniques. However, even with the help of automated synthesis and sequencing devices, many intermediate steps still require expert laboratory technicians to execute. We have developed an automated end-to-end DNA data storage device to explore the challenges of automation within the constraints of this unique application. Our device encodes data into a DNA sequence, which is then written to a DNA oligonucleotide using a custom DNA synthesizer, pooled for liquid storage, and read using a nanopore sequencer and a novel, minimal preparation protocol. We demonstrate an automated 5-byte write, store, and read cycle with a modular design enabling expansion as new technology becomes available.
阅读论文全文请访问:http://t.cn/E6sz5kk
期刊介绍:Scientific Reports (https://www.nature.com/srep/) is an online, open access journal from the publishers of Nature. We publish scientifically valid primary research from all areas of the natural and clinical sciences.
The 2017 journal metrics for Scientific Reports are as follows:
•2-year impact factor: 4.122
•5-year impact factor: 4.609
•Immediacy index: 0.576
•Eigenfactor® score: 0.71896
•Article Influence Score: 1.356
•2-year Median: 2
(来源:科学网)
特别声明:本文转载仅仅是出于传播信息的需要,并不意味着代表本网站观点或证实其内容的真实性;如其他媒体、网站或个人从本网站转载使用,须保留本网站注明的“来源”,并自负版权等法律责任;作者如果不希望被转载或者联系转载稿费等事宜,请与我们接洽。